DREAMS: deep read-level error model for sequencing data applied to
Por um escritor misterioso
Descrição
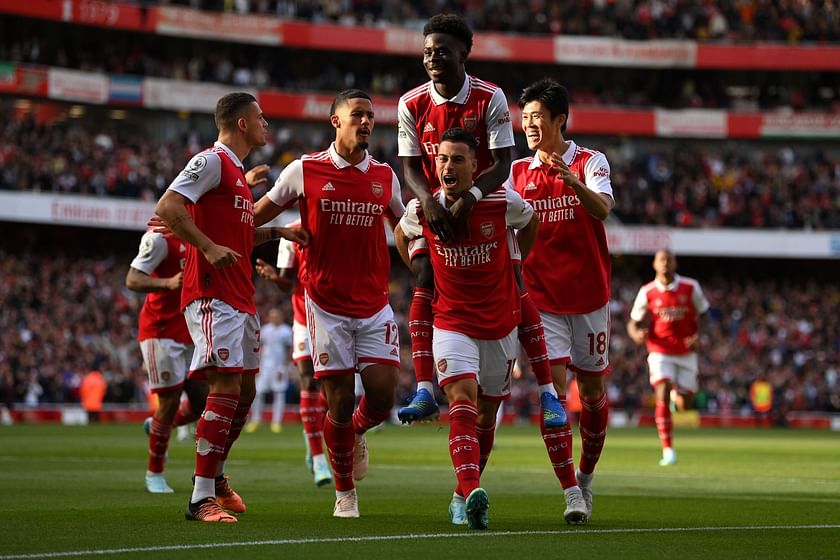
Circulating tumor DNA detection using next-generation sequencing (NGS) data of plasma DNA is promising for cancer identification and characterization. However, the tumor signal in the blood is often low and difficult to distinguish from errors. We present DREAMS (Deep Read-level Modelling of Sequencing-errors) for estimating error rates of individual read positions. Using DREAMS, we develop statistical methods for variant calling (DREAMS-vc) and cancer detection (DREAMS-cc). For evaluation, we generate deep targeted NGS data of matching tumor and plasma DNA from 85 colorectal cancer patients. The DREAMS approach performs better than state-of-the-art methods for variant calling and cancer detection.

Machine Learning in Predictive Toxicology: Recent Applications and

The Full Guide to Embeddings in Machine Learning
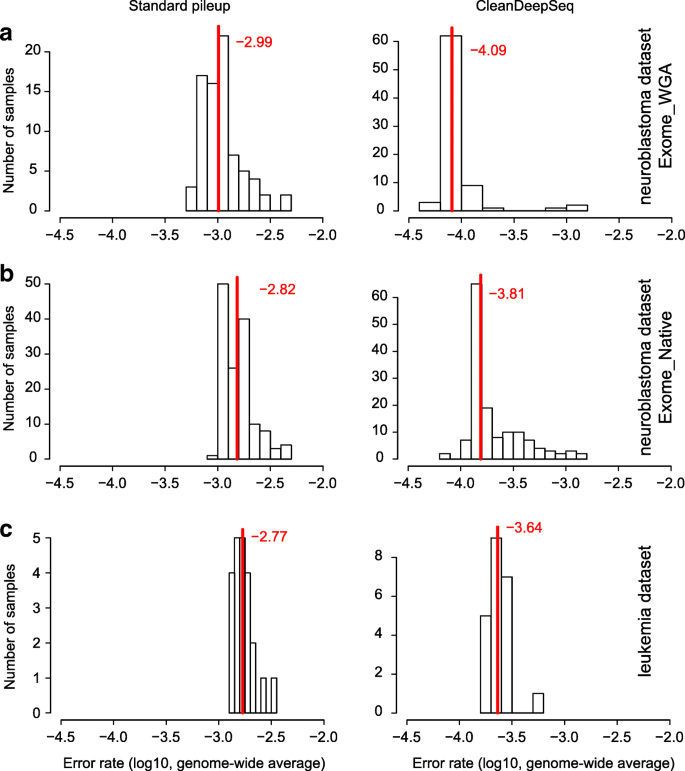
Analysis of error profiles in deep next-generation sequencing data

SpikingJelly: An open-source machine learning infrastructure
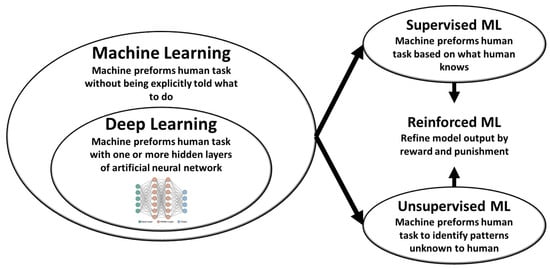
Life, Free Full-Text

DREAMS: Deep Read-level Error Model for Sequencing data applied to
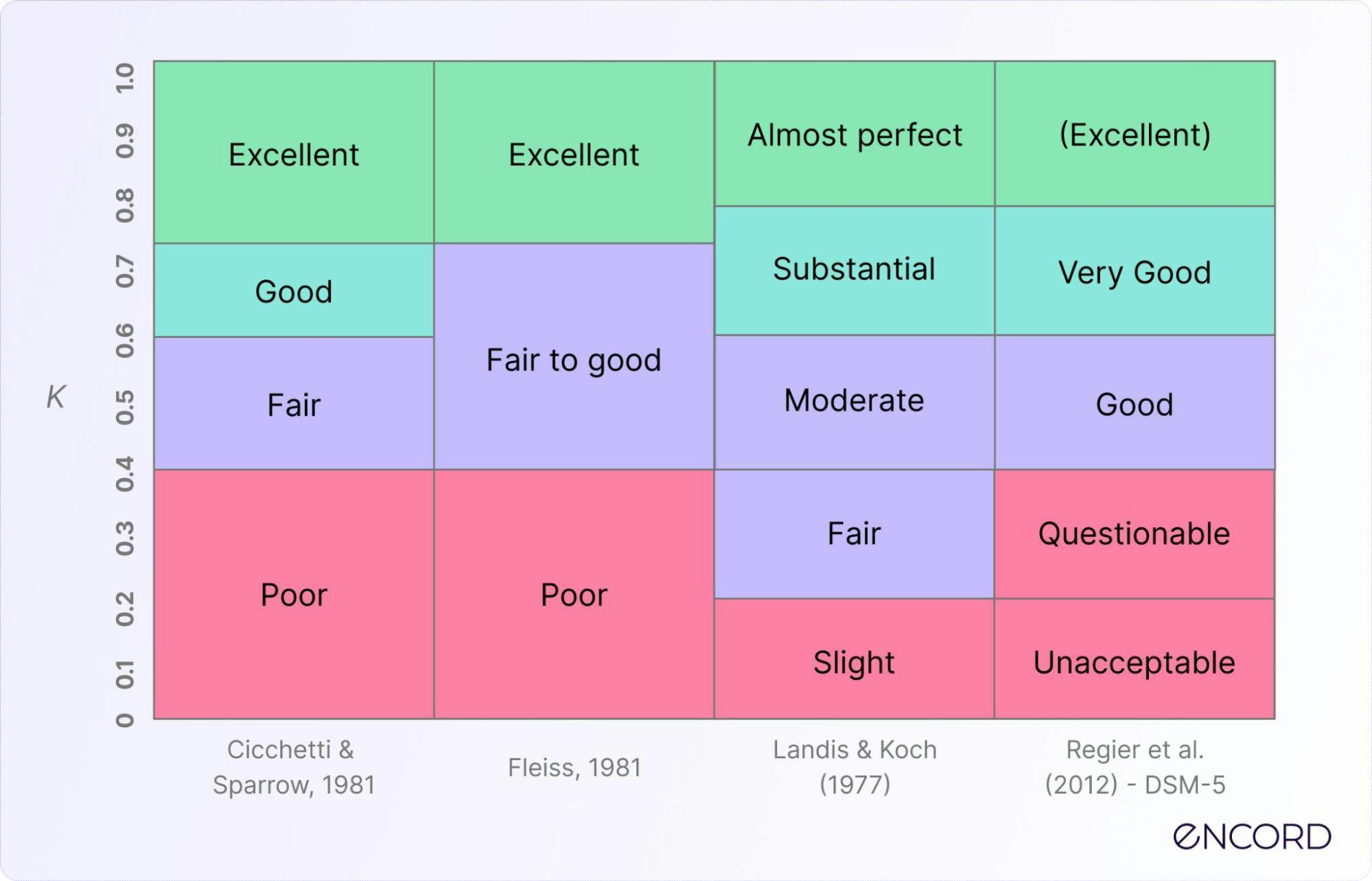
Inter-rater Reliability: Definition & Applications
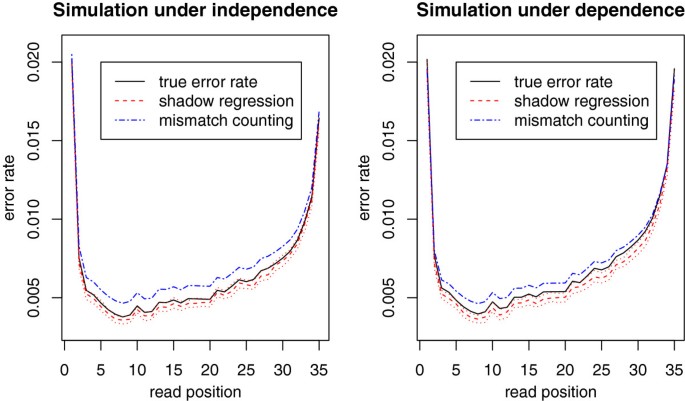
Estimation of sequencing error rates in short reads
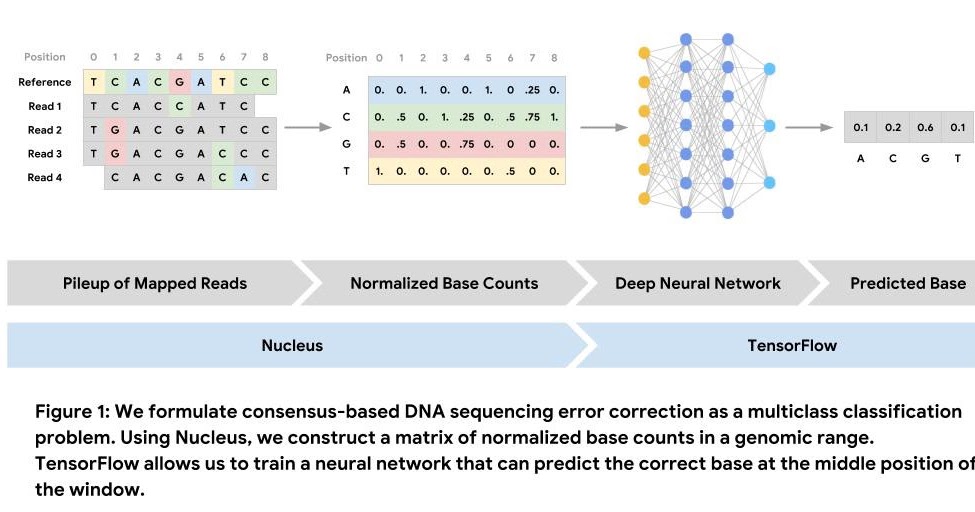
Using Nucleus and TensorFlow for DNA Sequencing Error Correction

DREAMS: Deep Read-level Error Model for Sequencing data applied to
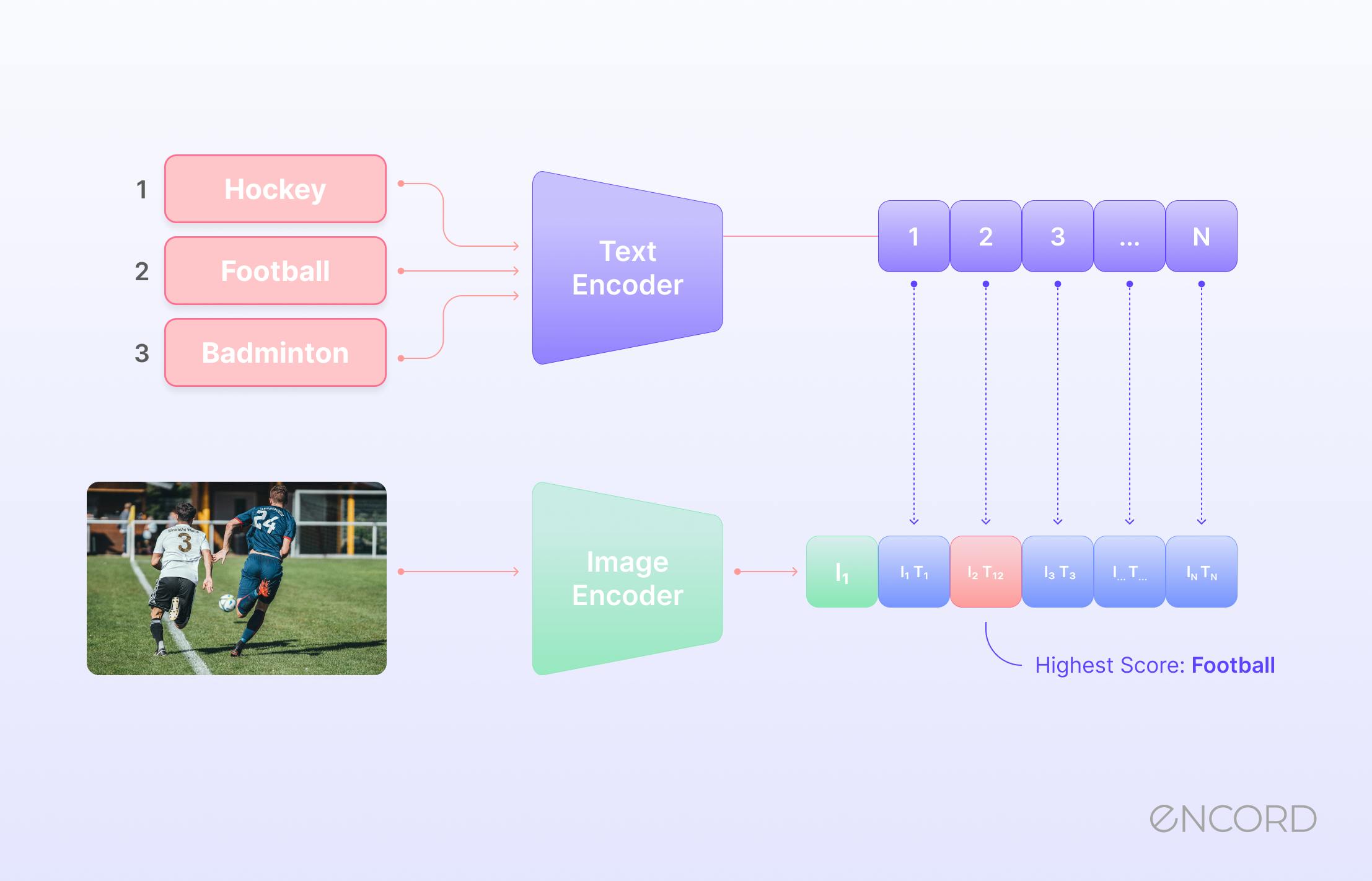
Zero-Shot Learning (ZSL) Explained: Applications, Challenges, and
de
por adulto (o preço varia de acordo com o tamanho do grupo)